Integration
Pipeline Integration
Pipeline Scripts
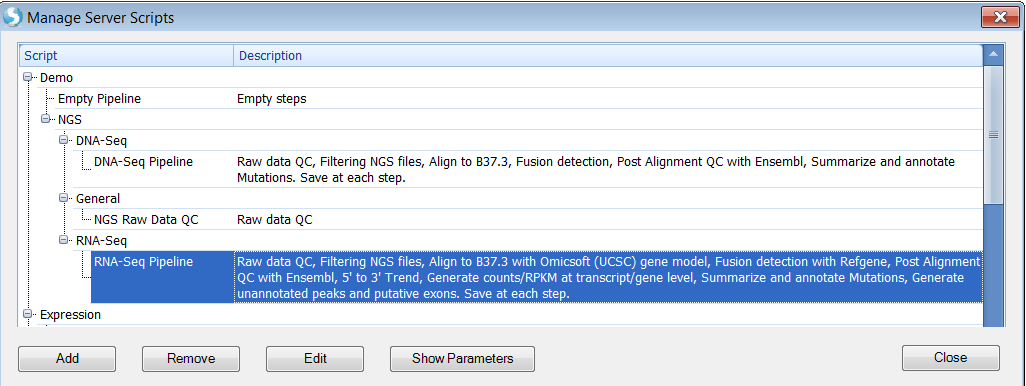
User can run a pre-configured pipeline on a sample set. Pipeline can be as simple as raw data QC, alignment and then do quantification analysis on NGS data. Pipeline scripts are managed by ArrayServer administrators in Server | Manage | Manage Scripts:
Run Pipeline Script in Analysis
To run a pipeline on a SampleSet, open or create a server project in the Analysis tab. Then go to Add Data | Add Data From Server:
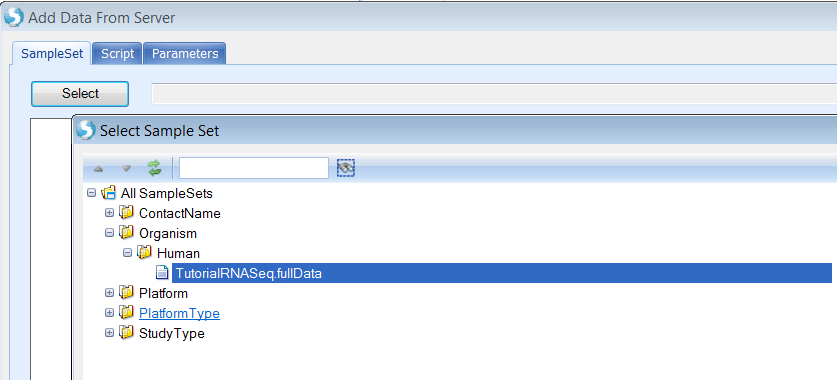
Select a SampleSet
Select a pipeline script:
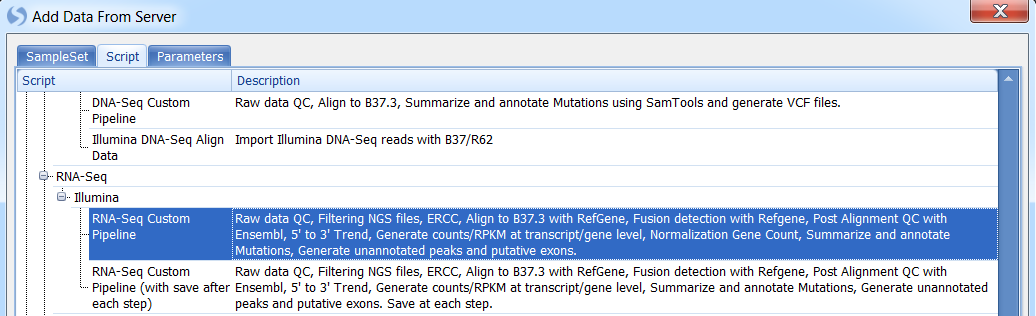
Fill required parameter values and click OK to run:
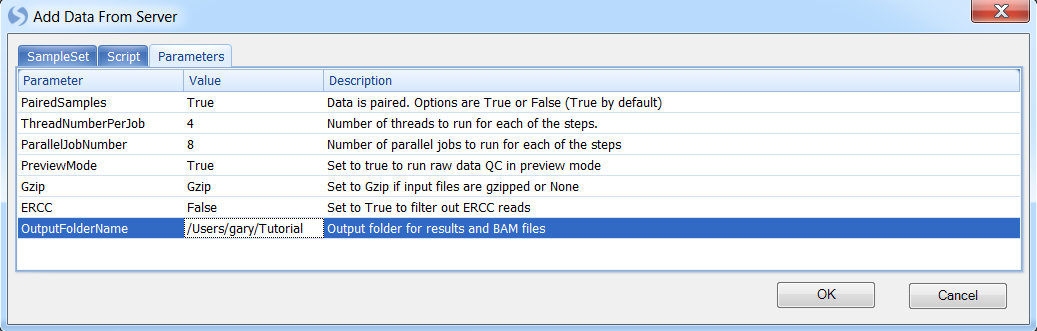
Run Pipeline Script during Sample Registration
User can run pipeline script during sample registration process by adding [Pipeline] section in sample registration file.
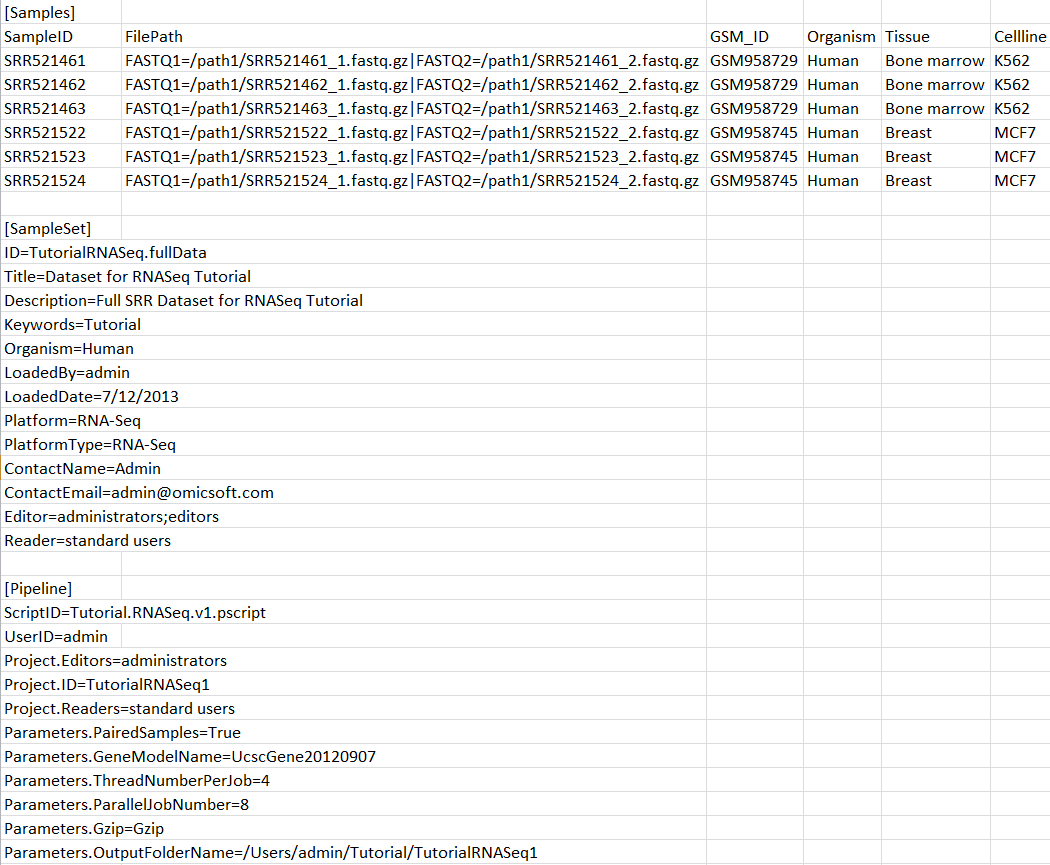
By using the sample registration file above, a server job will be submitted at the end of sample registration to create a server project with ID TutorialRNASeq1, using pipeline script Tutorial.RNASeq.v1.pscript and specified parameter values.
Genome Browser Integration
As we mentioned above, sample registration supports multiple file formats, including BAM (NGS alignment) files. User can modify the old registration file, adding a file path to BAM files:
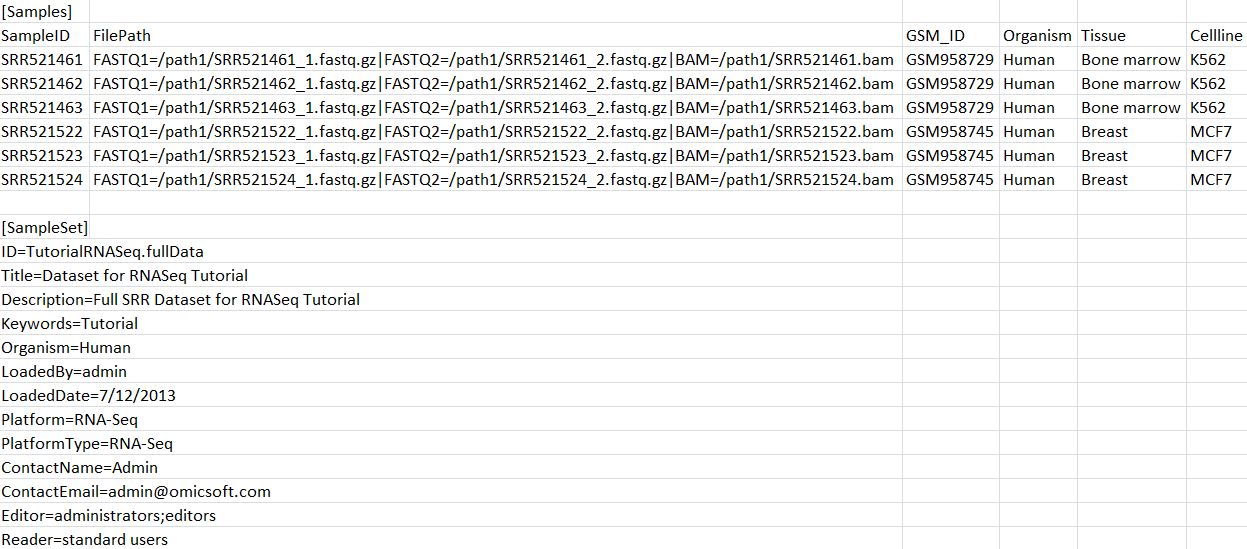
Registering this file (Server Sample | Register Samples) will update samples with BAM path.
Now, the alignment files (BAM files) of samples can be loaded in the Genome Browser tab directly through Add Track | Add Track From Server Samples:
User can add individual samples or the whole sample set. Here we choose the TutorialRNASeq.fullData sampleset:
User has the option to add one sample as one genome browser track:
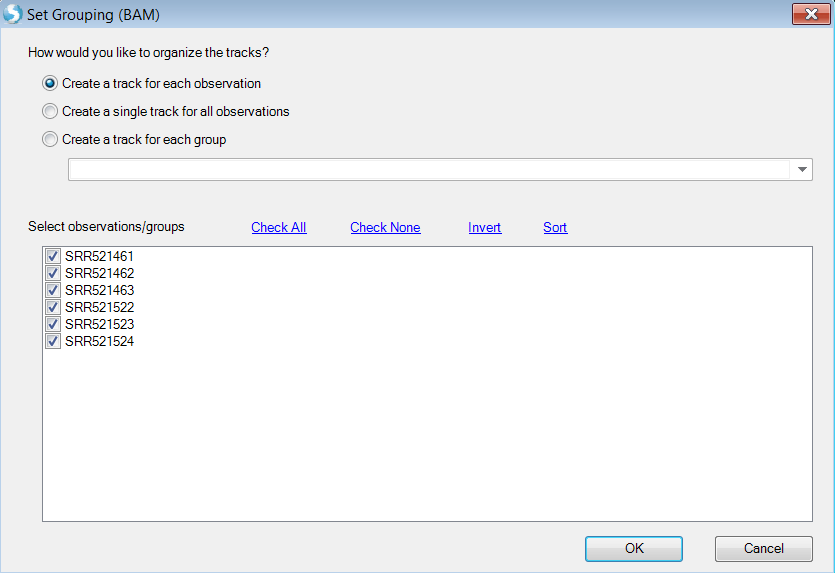
Or create tracks for groups defined by sample registration meta data:
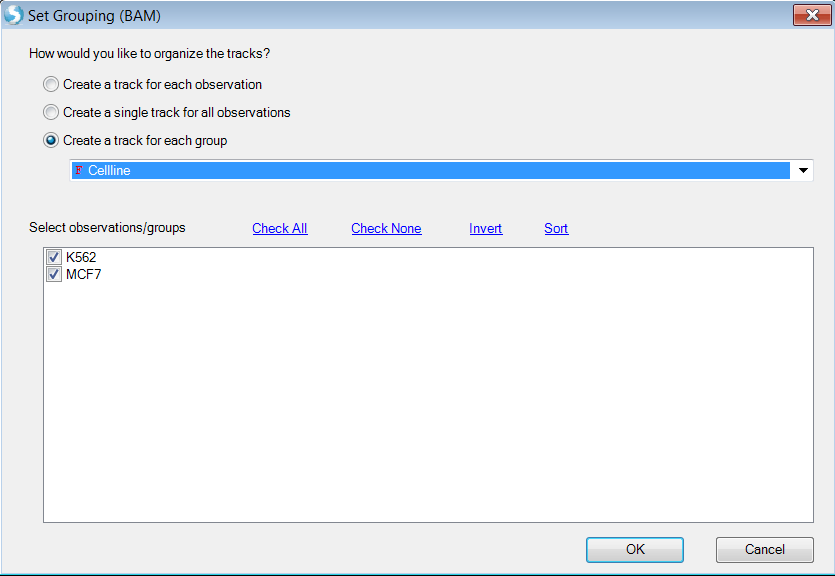
Choose Cellline as group which will create two genome browser tracks: K562 and MCF7.
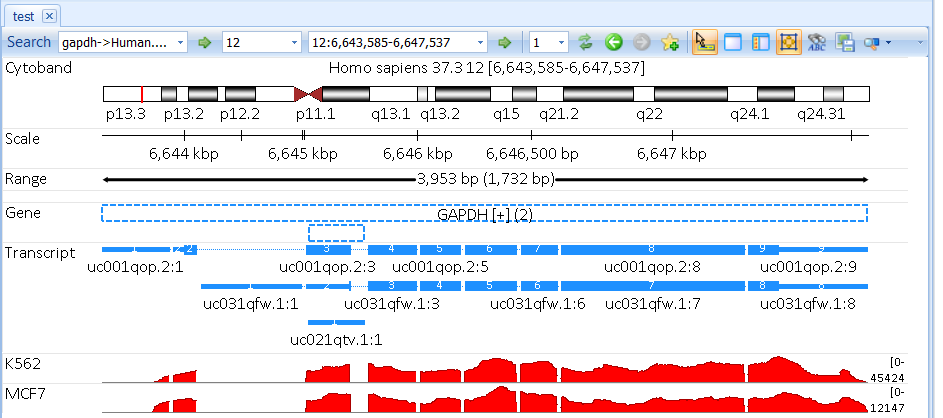
User can split the combined tracks to individual samples by right click and select Split Into Multiple Tracks:
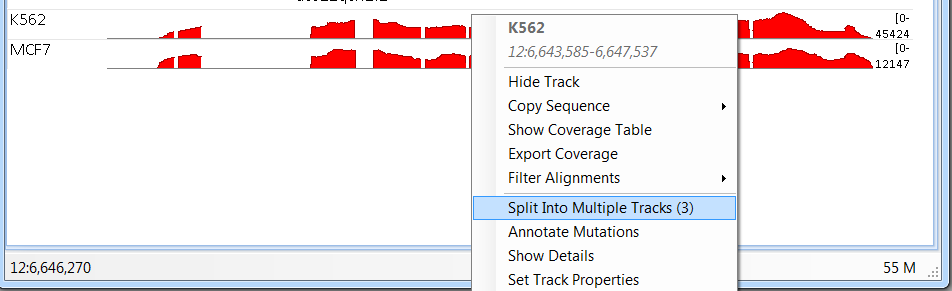
For more information and features available in the genome browser, please read the Genome Browser Tutorial.