Introduction
ArrayServer
Array Server is an enterprise solution, allowing users to store, share, search, and integrate their microarray/SNP/CNV/NGS projects and data. Easily share analyzed data with clients and colleagues. ArrayServer also hosts shared genome browsers. The following diagram demonstrates the functionality of Array Server:
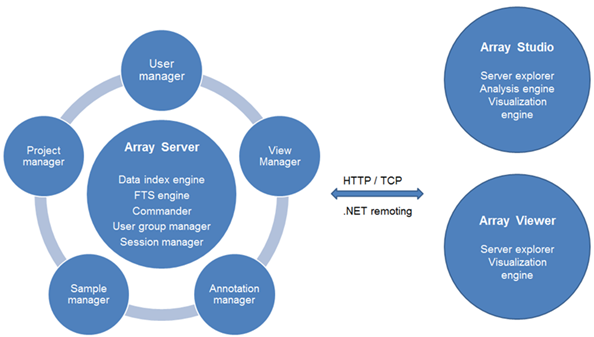
Server-based analysis allows the user to run jobs on a remote server (Linux or Windows), which usually has more computing power than a desktop computer. The Array Studio client software, installed on a local desktop machine, is used to interact with the Array Server (like a server terminal). Tasks such as job submission, monitoring, file transfer and data visualization can be done through the client software. Moreover, Array Server has a built-in scheduling system that supports high performance computing cluster (both SGE and PBS/Torque), accelerating the analysis of tremendous amounts of NGS/Omics data.
All the Array Studio modules available for local analysis are also accessible for server-based analysis. The graphic interfaces are almost the same.
Server Project
A "Server Project" is a project that is created on the server, rather than on the user's client machine. This project is cached locally on the client machine, in case of loss of connection to the server, but is automatically updated each time the user logs in or clicks the save button. The cache is stored in the user s home folder, typically My Documents/Omicsoft/ServerProjects. A Server Project, when stored locally in the cache folder, has a different filename suffix (.ossprj) compared to a regular project (.osprj) and can only be opened when the user is connected to the server.
The concept behind the Server Project is that any data that is added to a project must first be stored on the server. When the user adds a new dataset (whether it be Gene Expression, CNV, or NextGen sequencing), they will be prompted with the folder structure of the Array Server instead of their local file system. If the user wishes to use a file from their local file system, they must first upload the file to the server file system. Most of companies are storing data in network drives, they can map these drives directly in Array Server and all users will be able to access data easily during server analysis.
All data addition and extraction is done on the server side, by Array Server, instead of the users client machine. It allows the user to use the power of the server, instead of their individual client machine for the importing of data. This is extremely important for some memory/CPU intensive importing operations (such as alignment of a fastq file to the genome for NextGen sequencing data). After data extraction, alignment or data summarization, data can be downloaded to the client machine, where the user can add views, run additional statistical analysis, etc. Anytime the user saves the project, it is synchronized with the version on the server.
Test Dataset
In this tutorial, we will show how to create a server project for server-based analysis. The user should first download the demo data file for use in the tutorial and save it to the local machine. The file can be accessed from the following URL: link